Volumes (DICOM)
Introduction
How to debug this tutorial
Import libraries
Init API client
Get variables from environment
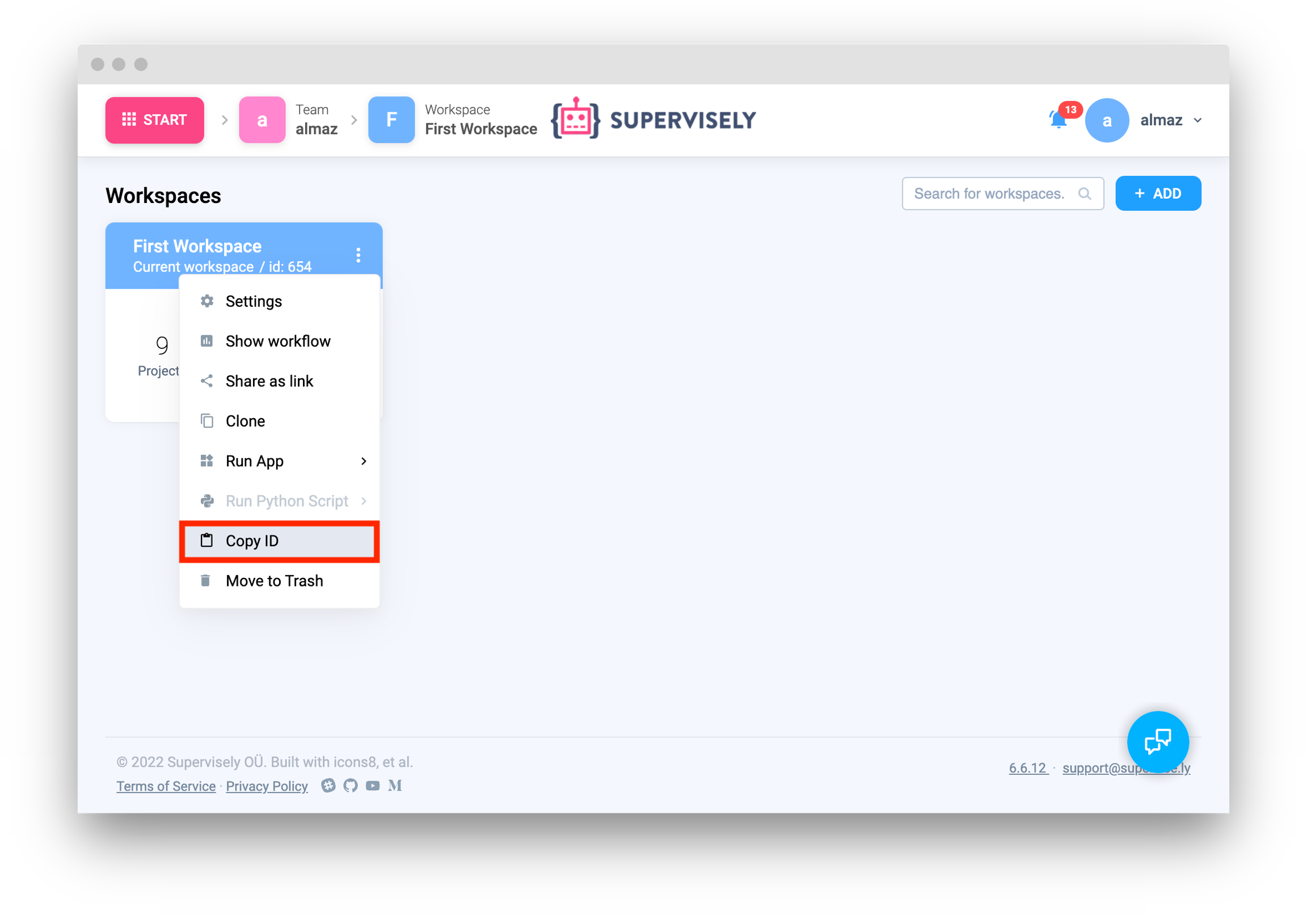
Create new project and dataset
Upload volumes from local directory to Supervisely
Upload NRRD format volume
Upload volume as NumPy array
Upload DICOM series from local directory
Upload list of volumes from local directory
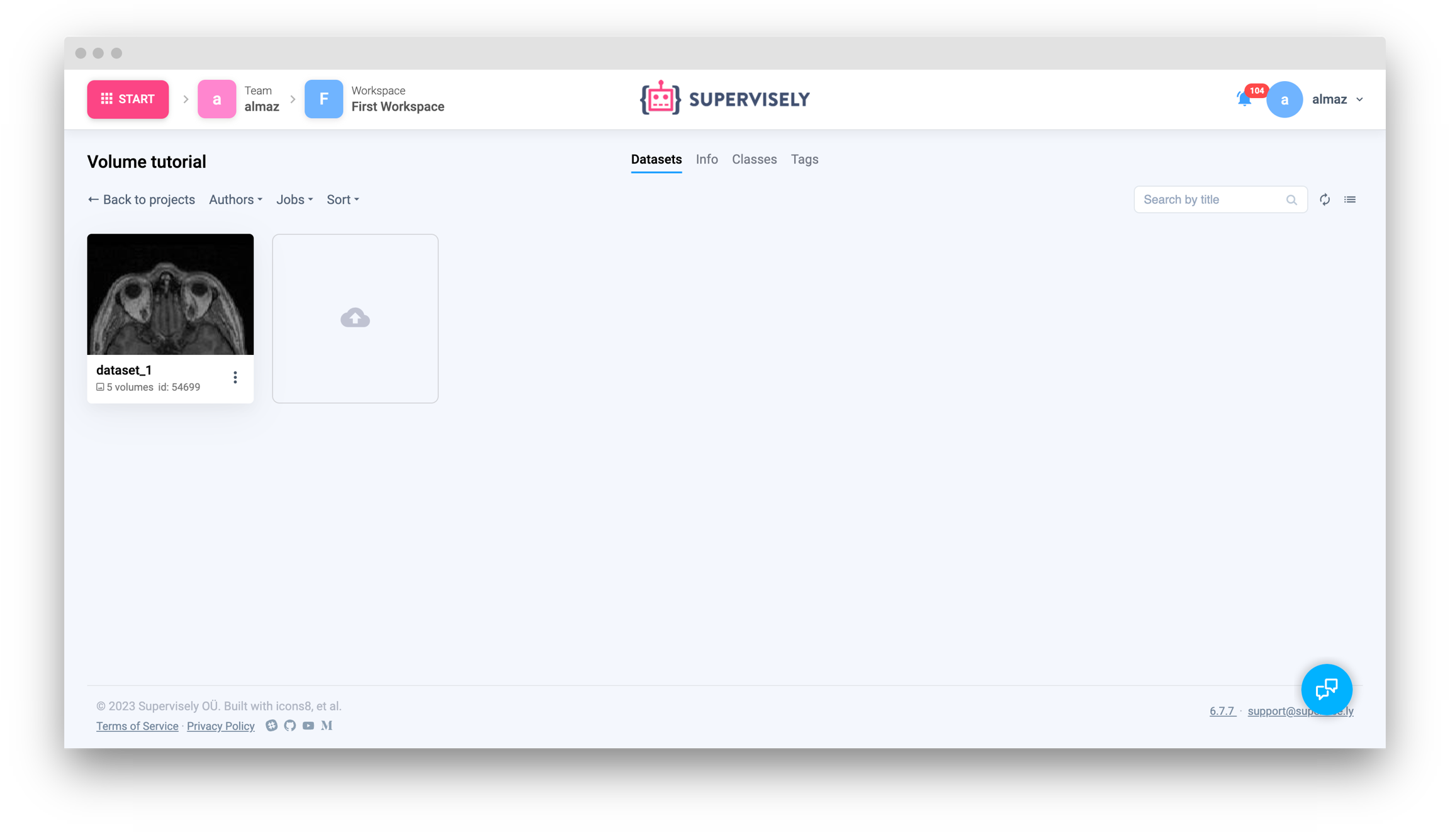
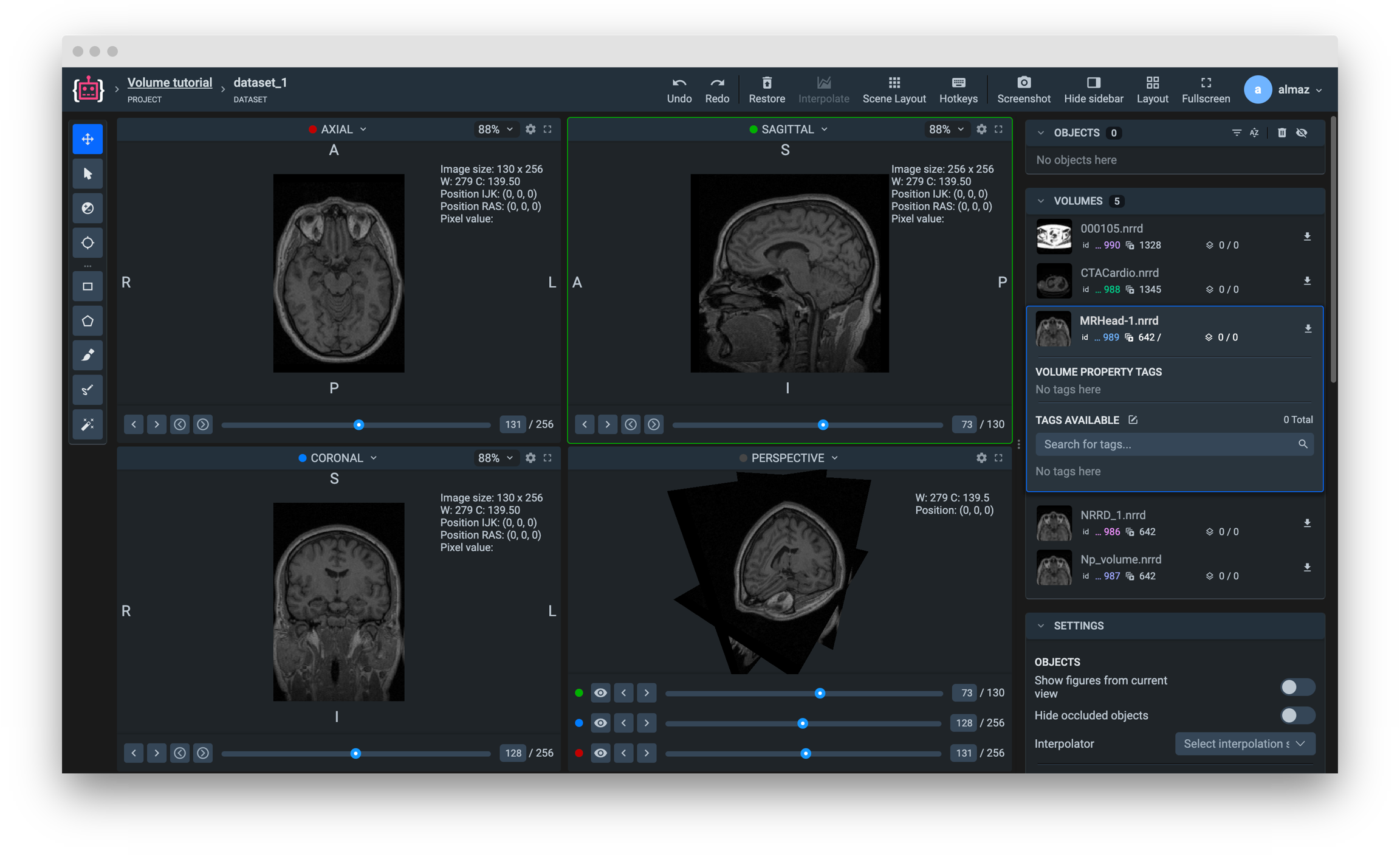
Get volume info from Supervisely
Get list of volumes infos from current dataset
Get single volume info by id
Get single volume info by name
Download volume from Supervisely to local directory
Get volume slices from local directory
Read NRRD file from local directory
Get slices from volume
Download slice from Supervisely
Save slice to local directory
Save slice as NRRD
Save slice as JPG
Note:
Last updated